Folks know that gradient-boosted trees generally perform better than a random forest, although there is a price for that: GBT have a few hyperparams to tune, while random forest is practically tuning-free. Let’s look at what the literature says about how these two methods compare.
Supervised learning in 2005
In 2005, Caruana et al. made an empirical comparison of supervised learning algorithms [video]. They included random forests and boosted decision trees and concluded that
With excellent performance on all eight metrics, calibrated boosted trees were the best learning algorithm overall. Random forests are close second.
Let’s note two things here. First, they mention calibrated boosted trees, meaning that for probabilistic classification trees needed calibration to be the best. Second, it’s unclear what boosting method the authors used.
In the follow-up study concerning supervised learning in high dimensions the results are similar:
Although there is substantial variability in performance across problems and metrics in our experiments, we can discern several interesting results. First, the results confirm the experiments in (Caruana & Niculescu-Mizil, 2006) where boosted decision trees perform exceptionally well when dimensionality is low. In this study boosted trees are the method of choice for up to about 4000 dimensions. Above that, random forests have the best overall performance.
Hundreds of classifiers
Ten years later Fernandez-Delgado et al. revisited the topic with the paper titled Do we need hundreds of classifiers to solve real world classification problems? Notably, there were no results for gradient-boosted trees, so we asked the author about it. Here’s the answer, reprinted with permission:
That comment has been issued by other researcher (David Herrington), our response was that we tried GBM (gradient boosting machine) in R directly and via caret, but we achieved errors for problems with more than two[-class] data sets. However, in response to him, we developed further experiments with GBM (using only two-class data sets) achieving good results, even better than random forest but only for two-class data sets. This is the email with the results. I hope they can be useful for you. Best regards!
The email
Dear Prof. Herrington:
I apologize for the delay in the answer to your last email. I have achieved results using gbm, but I was so delayed because I found errors with data sets more than two classes: gbm with caret only worked with two-class data sets, it gives an error with multi-class data sets, the same error as in http://stackoverflow.com/questions/15585501/usage-of-caret-with-gbm-method-for-multiclass-classification.
I tried to run gbm directly in R as tells the previous link, but I also found errors with multi-class data sets. I have been trying to find a program that runs, but I did not get it. I will keep trying, but by now I send to you the results with two classes, comparing both GBM and Random Forests (in caret, i.e., rf_t in the paper). The GBM worked without only for 51 data sets (most of them with two classes, although there are 55 data sets with two classes, so that GBM gave errors in 4 two-class data sets), and the average accuracies are:
rf = 82.30% (+/-15.3), gbm = 83.17% (+/-12.5)
so that GBM is better than rf_t. In the paper, the best classifier for two-class data sets was avNNet_t, with 83.0% accuracy, so that GBM is better on these 51 data sets. Attached I send to you the results of RF and GBM, and the plot with the two accuracies (ordered decreasingly) for the 51 data sets.
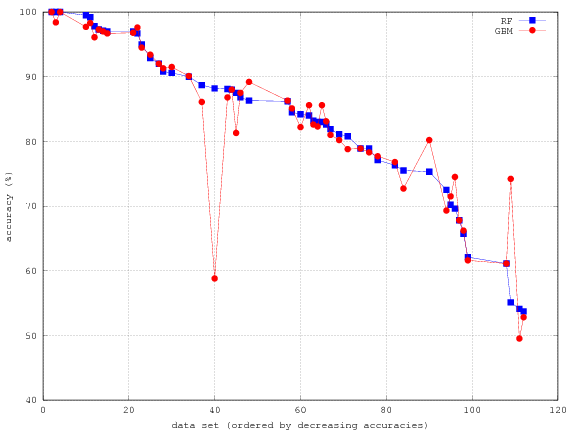
The detailed results are available on GitHub.
Tuning
From the chart it would seem that RF and GBM are very much on par. Our feeling is that GBM offers a bigger edge. For example, in Kaggle competitions XGBoost replaced random forests as a method of choice (where applicable).
If we were to guess, the edge didn’t show in the paper because GBT need way more tuning than random forests. It’s quite time consuming to tune an algorithm to the max for each of the many datasets.
Random forest
With a random forest, in contrast, the first parameter to select is the number of trees. Easy: the more, the better. That’s because the multitude of trees serves to reduce variance. Each tree fits, or overfits, a part of the training set, and in the end their errors cancel out, at least partially. Random forests do overfit, just compare the error on train and validation sets.
Other parameters you may want to look at are those controlling how big a tree can grow. As mentioned above, averaging predictions from each tree counteracts overfitting, so usually one wants biggish trees.
One such parameter is min. samples per leaf. In scikit-learn’s RF, it’s value is one by default. Sometimes you can try increasing this value a little bit to get smaller trees and less overfitting. This CoverType benchmark overdoes it, going from 1 to 13 at once. Try 2 or 3 first.
Finally, there’s max. features to consider. Once upon a time, we tried tuning that param, to no avail. We suspect that it may have a better effect when dealing with sparse data - it would make sense to try increasing it then.
That’s about it for random forests. With gradient-boosted trees there are so many parameters that it’s a subject for a separate article.